 LEVEL2 is an implementation in Java of the algorithm
of Leo van Iersel, Judith Keijsper, Steven Kelk and Leen Stougie
which constructs level-2 phylogenetic networks from rooted
triplets. The code was written, and is maintained, by
Steven Kelk.
LEVEL2 is an implementation in Java of the algorithm
of Leo van Iersel, Judith Keijsper, Steven Kelk and Leen Stougie
which constructs level-2 phylogenetic networks from rooted
triplets. The code was written, and is maintained, by
Steven Kelk.
 LEVEL2 is an implementation in Java of the algorithm
of Leo van Iersel, Judith Keijsper, Steven Kelk and Leen Stougie
which constructs level-2 phylogenetic networks from rooted
triplets. The code was written, and is maintained, by
Steven Kelk.
LEVEL2 is an implementation in Java of the algorithm
of Leo van Iersel, Judith Keijsper, Steven Kelk and Leen Stougie
which constructs level-2 phylogenetic networks from rooted
triplets. The code was written, and is maintained, by
Steven Kelk.
NOTE: As of January 28th 2009 the latest release of LEVEL2 will be available from Sourceforge under a GNU Public Licence. Please follow this link for the most recent release. There you can download the Java source and all the most recent documentation and examples.
The following instructions assume a UNIX environment. There are no fundamental reasons however why the algorithm will not work under another operating system environment as long as Java and the graph drawing package DOT are available in that environment. (Please contact Steven Kelk if you are interested in an output format for a package other than DOT.)
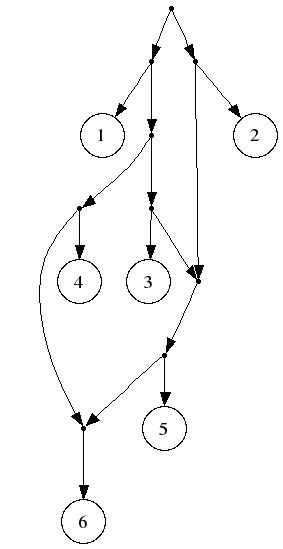
From real yeast data, obtained from the Fungal Biodiversity Center in Utrecht (Netherlands), the following triplets were generated, producing the following 21-leaf level-2 network (scalable EPS version is here.) Biological analysis of this data will appear in a future publication.
Some more examples can be found in this directory. Note that the networks generated by LEVEL2 are always planar, but that sometimes DOT will draw the network in non-planar fashion. This is something which will hopefully be addressed in a future version.
For a detailed analysis of the algorithm, please consult our article. For a summary of the algorithm see the following powerpoint presentation (summer 2007).